I’ve moved the Pyevolve SVN repository hosted at sourceforge.net to a new Github repository.
The new official Pyevolve repository is now located at https://github.com/perone/Pyevolve.
by Christian S. Perone
I’ve moved the Pyevolve SVN repository hosted at sourceforge.net to a new Github repository.
The new official Pyevolve repository is now located at https://github.com/perone/Pyevolve.
I would like to share the video of the Eric Floehr (from ForecastWatch / Intellovations) presentation at PyCon 2011:
Eric Floehr (from Intellovations) kindly sent me the presentation he presented at PyOhio 2010. I think Eric has captured some nice features of Pyevolve which few people use, like DB Adapters, dot plotting, Interactive Mode, Real Time statistics, etc. He also presents an interesting use case where he uses Genetic Programming in order to forecast weather based on some historical data:
Following the recent article arguing why PyPy is the future of Python, I must say, PyPy is not the future of Python, is the present. When I have tested it last time (PyPy-c 1.1.0) with Pyevolve into the optimization of a simple Sphere function, it was at least 2x slower than Unladen Swallow Q2, but in that time, PyPy was not able to JIT. Now, with this new release of PyPy and the JIT’ing support, the scenario has changed.
PyPy has evolved a lot (actually, you can see this evolution here), a nice work was done on the GC system, saving (when compared to CPython) 8 bytes per object allocated, which is very interesting for applications that makes heavy use of object allocation (GP system are a strong example of this, since when they are implemented on object oriented languages, each syntax tree node is an object). Efforts are also being done to improve support for CPython extensions (written in C/C++), one of them is a little tricky: the use of RPyC, to proxy through TCP the remote calls to CPython; but the other seems by far more effective, which is the creation of the CPyExt subsystem. By using CPyExt, all you need is to have your CPython API functions implemented in CPyExt, a lot of people is working on this right now and you can do it too, it’s a long road to have a good API coverage, but when you think about advantages, this road becomes small.
In order to benchmark CPython, Jython, CPython+Psyco, Unladen Swallow and PyPy, I’ve used the Rastrigin function optimization (an example of that implementation is here in the Example 7 of Pyevolve 0.6rc1):
Due to its large search space and number of local minima, Rastrigin function is often used to measure the performance of Genetic Algorithms. Rastrigin function has a global minimum at where the
; in order to increase the search space and required resources, I’ve used 40 variables (
) and 10k generations.
Here are the information about versions used in this benchmark:
No warmup was performed in JVM or in PyPy. PyPy translator was executed using the “-Ojit” option in order to get the JIT version of the Python interpreter. The JVM was executed using the server mode, I’ve tested the client and server mode for Sun JVM and IcedTea6, the best results were observed from the server mode using Sun JVM, however when I’ve compared the client mode of IcedTea6 with the client mode of Sun JVM, the best results observed were from IcedTea6 (the same as using server mode in IcedTea6). Unladen Swallow was compiled using the project wiki instructions for building an optimized binary.
The machine used was an Intel(R) Core(TM) 2 Duo E4500 (2×2.20Ghz) with 2GB of RAM.
The result of the benchmark (measured using wall time) in seconds for each setup (these results are the best of 3 sequential runs):
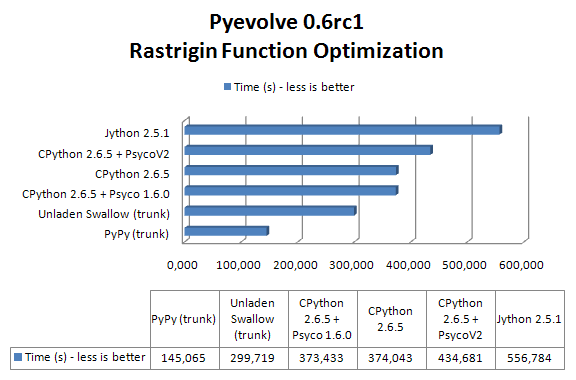
As you can see, PyPy with JIT got a speedup of 2.57x when compared to CPython 2.6.5 and 2.0x faster than Unladen Swallow current trunk.
PyPy is not only the future of Python, but is becoming the present right now. PyPy will not bring us only an implementation of Python in Python (which in itself is the valuable result of great efforts), but also will bring the performance back (which many doubted at the beginning, wondering how could it be possible for an implementation of Python in Python be faster than an implementation in C ? And here is where the translation and JIT magic enters). When the time comes that Python interpreter can be entire written in a high level language (actually almost the same language, which is really weird), Python community can put their focus on improving the language itself instead of spending time solving the complexity of the lower level languages, is this not the great point of those efforts ?
By the way, just to note, PyPy isn’t only a translator for the Python interpreter written in RPython, it’s a translator of RPython, what means that PyPy isn’t only the future of Python, but probably, the future of many interpreters.
I’m proud to announce the Pyevolve 0.6rc1 ! This is the first release candidate, but it’s pretty stable for production use (as from this 0.6 version we are very closer to a stable codebase, thanks to community).
See the documentation site and the What’s New.
I would like to thank the people who directly or indirectly have contributed with this release: Boris Gorelik, Amit Saha, Jelle Feringa, Henrik Rudstrom, Matteo de Felice, SIGEVOlution Board, Mike Benoit, Ryan Campbell, Jonas A. Gustavsson, Soham Sadhu, Ernesto Costa, Ido Ben-Tsion, Frank Goodman, Vishal Belsare, Benjamin Bush; and a lot of people who gave us feedback of the experience with the use of Pyevolve on their applications.
For downloads, go to the Downloads section at Documentation site.
If you see something wrong with this release candidate, please create a ticket reporting your problem at Pyevolve Trac, so we can provide fixes to the official release.
Happy coding !
– Christian S. Perone
Here is the video of Pyevolve (the development version) optimizing a 100 cities TSP problem. I’ve used the Edge Recombination and the simple Swap Mutation method:
The video is a composition the of image outputs at every 100th generation and only when the score has changed when compared to the previous generation. I’ll release the source together with the new 0.6 release, planned to this month (December) and no later than January.
Best wishes and if I can’t post again until the next year, merry christmas and happy new year !!!
– Christian S. Perone
I’m proud to announce that Pyevolve is featuring on the last issue of SIGEVOlution (Volume 4, Issue 1), a newsletter from the ACM Special Interest Group on Evolutionary Computation. I would like to thank the newsletter editor Pier Luca Lanzi and the board for the corrections in the article and for the well done reformatted version of the paper.
Pyevolve is currently in version 0.5, in a few months I’ll be releasing the new 0.6 release with the new major features that are currently implemented in the development version only (you can check it at the subversion repository in sourceforge.net).
I hope you enjoy the article !
Yours,
– Christian S. Perone
So I did a comparative of Pyevolve GP/GA core in different Python interpreters. I’ve used my Pentium Core 2 Duo (E4500 @ 2.20GHz, 1GB RAM), using Ubuntu 9.04 and Windows XP SP3 just for IronPython 2.6.1 (IronPython doesn’t run with Mono, so I used the win xp with .net 2.0).
The interpreters used were:
I tried using 2009Q3 (the currently main trunk), but I think it’s unstable yet, cause it was more slow than 2009Q2, so I used 2009Q2; I compiled it with GCC 4.3.3 just using the default configure parameters (./configure).
I used the default CPython package of Ubuntu 9.04.
I used the default CPython package of Ubuntu 9.04 too, the python2.5 package.
I used the last svn version of the repository, the release 68612. My Pentium Core 2 Duo had only 1GB of RAM, and the PyPy translation process eats more RAM than Java (sorry for the joke), so I used a notebook with 3GB of RAM to create the pypy-c, what took 1 hour (I used –opt=3) and a beautiful ascii Mandelbrot fractal !
I used the default installer from the Jython project site. I used the Sun JRE 1.6.0_16.
I’ve used the 2.6 RC1 available at IronPython project site with MS .NET 2.0.
To test the GA core I’ve used this source-code (a simple sphere function):
from pyevolve import G1DList
from pyevolve import Mutators, Initializators
from pyevolve import GSimpleGA, Consts
# This is the Sphere Function
def sphere(xlist):
total = 0
for i in xlist:
total += i**2
return total
def run_main():
genome = G1DList.G1DList(140)
genome.setParams(rangemin=-5.12, rangemax=5.13)
genome.initializator.set(Initializators.G1DListInitializatorReal)
genome.mutator.set(Mutators.G1DListMutatorRealGaussian)
genome.evaluator.set(sphere)
ga = GSimpleGA.GSimpleGA(genome, seed=666)
ga.setMinimax(Consts.minimaxType["minimize"])
ga.setGenerations(1500)
ga.setMutationRate(0.01)
ga.evolve(freq_stats=500)
best = ga.bestIndividual()
if __name__ == "__main__":
run_main()
And to test the GP core, I’ve used this source-code (a simple symbolic regression):
from pyevolve import GTree
from pyevolve import Mutators
from pyevolve import GSimpleGA, Consts, Util
import math
rmse_accum = Util.ErrorAccumulator()
def gp_add(a, b): return a+b
def gp_sub(a, b): return a-b
def gp_mul(a, b): return a*b
def gp_sqrt(a): return math.sqrt(abs(a))
def eval_func(chromosome):
global rmse_accum
rmse_accum.reset()
code_comp = chromosome.getCompiledCode()
for a in xrange(0, 10):
for b in xrange(0, 10):
evaluated = eval(code_comp)
target = math.sqrt((a*a)+(b*b))
rmse_accum += (target, evaluated)
return rmse_accum.getRMSE()
def main_run():
genome = GTree.GTreeGP()
genome.setParams(max_depth=4, method="ramped")
genome.evaluator += eval_func
genome.mutator.set(Mutators.GTreeGPMutatorSubtree)
ga = GSimpleGA.GSimpleGA(genome, seed=666)
ga.setParams(gp_terminals = ['a', 'b'],
gp_function_prefix = "gp")
ga.setMinimax(Consts.minimaxType["minimize"])
ga.setGenerations(40)
ga.setCrossoverRate(1.0)
ga.setMutationRate(0.08)
ga.setPopulationSize(800)
ga(freq_stats=10)
best = ga.bestIndividual()
if __name__ == "__main__":
main_run()
UPDATE 19/08: the x-axis is measured in “seconds“, and the y-axis is the python flavor;
The results are are described in the graph below:
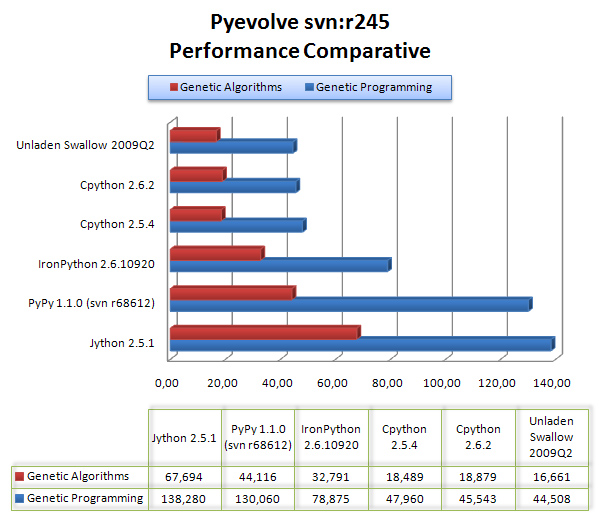 As we can see, Unladen Swallow 2009Q2 did a little better performance than CPython 2.6.2, but Jython and PyPy (experimental) were left behind in that scenario, even behind IronPython 2.6.1.
As we can see, Unladen Swallow 2009Q2 did a little better performance than CPython 2.6.2, but Jython and PyPy (experimental) were left behind in that scenario, even behind IronPython 2.6.1.