The future can be written in RPython now
Following the recent article arguing why PyPy is the future of Python, I must say, PyPy is not the future of Python, is the present. When I have tested it last time (PyPy-c 1.1.0) with Pyevolve into the optimization of a simple Sphere function, it was at least 2x slower than Unladen Swallow Q2, but in that time, PyPy was not able to JIT. Now, with this new release of PyPy and the JIT’ing support, the scenario has changed.
PyPy has evolved a lot (actually, you can see this evolution here), a nice work was done on the GC system, saving (when compared to CPython) 8 bytes per object allocated, which is very interesting for applications that makes heavy use of object allocation (GP system are a strong example of this, since when they are implemented on object oriented languages, each syntax tree node is an object). Efforts are also being done to improve support for CPython extensions (written in C/C++), one of them is a little tricky: the use of RPyC, to proxy through TCP the remote calls to CPython; but the other seems by far more effective, which is the creation of the CPyExt subsystem. By using CPyExt, all you need is to have your CPython API functions implemented in CPyExt, a lot of people is working on this right now and you can do it too, it’s a long road to have a good API coverage, but when you think about advantages, this road becomes small.
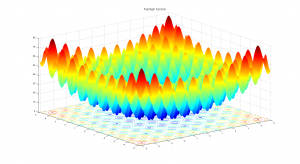
In order to benchmark CPython, Jython, CPython+Psyco, Unladen Swallow and PyPy, I’ve used the Rastrigin function optimization (an example of that implementation is here in the Example 7 of Pyevolve 0.6rc1):
Due to its large search space and number of local minima, Rastrigin function is often used to measure the performance of Genetic Algorithms. Rastrigin function has a global minimum at where the
; in order to increase the search space and required resources, I’ve used 40 variables (
) and 10k generations.
Here are the information about versions used in this benchmark:
- OS Ubuntu Linux 10.04 LTS (lucid)
- CPython 2.6.5 (Apr 16 2010)
- Unladen Swallow trunk (r1153)
- Jython 2.5.1 (Sun JVM 1.6.0_20, Server Mode)
- CPython 2.6.5 + PsycoV2 trunk (r74587)
- CPython 2.6.5 + Psyco 1.6.0 (default lucid package)
- PyPy trunk (r74537)
No warmup was performed in JVM or in PyPy. PyPy translator was executed using the “-Ojit” option in order to get the JIT version of the Python interpreter. The JVM was executed using the server mode, I’ve tested the client and server mode for Sun JVM and IcedTea6, the best results were observed from the server mode using Sun JVM, however when I’ve compared the client mode of IcedTea6 with the client mode of Sun JVM, the best results observed were from IcedTea6 (the same as using server mode in IcedTea6). Unladen Swallow was compiled using the project wiki instructions for building an optimized binary.
The machine used was an Intel(R) Core(TM) 2 Duo E4500 (2×2.20Ghz) with 2GB of RAM.
The result of the benchmark (measured using wall time) in seconds for each setup (these results are the best of 3 sequential runs):
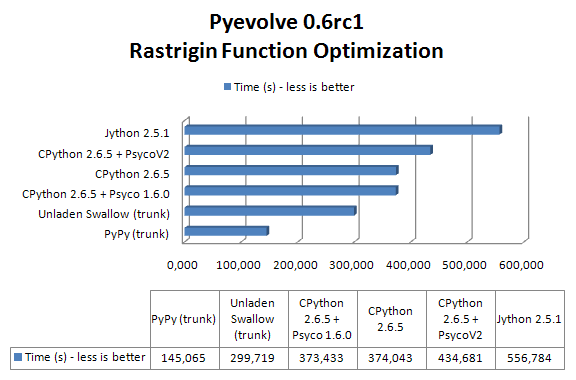
As you can see, PyPy with JIT got a speedup of 2.57x when compared to CPython 2.6.5 and 2.0x faster than Unladen Swallow current trunk.
PyPy is not only the future of Python, but is becoming the present right now. PyPy will not bring us only an implementation of Python in Python (which in itself is the valuable result of great efforts), but also will bring the performance back (which many doubted at the beginning, wondering how could it be possible for an implementation of Python in Python be faster than an implementation in C ? And here is where the translation and JIT magic enters). When the time comes that Python interpreter can be entire written in a high level language (actually almost the same language, which is really weird), Python community can put their focus on improving the language itself instead of spending time solving the complexity of the lower level languages, is this not the great point of those efforts ?
By the way, just to note, PyPy isn’t only a translator for the Python interpreter written in RPython, it’s a translator of RPython, what means that PyPy isn’t only the future of Python, but probably, the future of many interpreters.

Nice!
Very exciting results. Any idea of what the comparative memory usage is like?
I will make another benchmark soon I get more time, to check memory footprint between implementations (using GP this time, which is memory intensive). Thanks for commenting =)
This is kinda weird to see Jython performing so badly. Why didn’t you include warmup rounds for Jython? It’s known that startup time of JVM is pretty big and warmups are required to have adequate benchmarks(let HotSpot do it’s job in JVM).
In my opinion, the start-up time makes part of the whole run, would be nice to have it cited, but it’s part of the whole run since you’ll have to lose this time anyway (specially in optimization problems for instance). If you run a test with the other interpreters and with Jython, you’ll see that the startup time is amortized in the long run time, what makes Jython so slow in the benchmarks isn’t the the start-up time, but the overall performance, even when the server mode is enabled, you can note this in the course of the evolution, the time required to evolve 500 generations for example, is very high when compared with other interpreters (and even after the jvm has performed JIT optimizations like inlining, etc…).
Could you please time a C implementation of the function optimization also? It would be a nice reference point.
How about adding a comparison with Cython?
Hm.
I’m getting increasingly annoyed by cython PR. Let’s face it – Cython is *not* a python runtime and as such doesn’t have to be included in every python runtimes comparison whether developers like it or not. It can be included, but it’s not mandatory, especially if module X does not compile under cython
I agree that the comparison wouldn’t be fair, but a Cython implementation would still be the easiest way to get a good base line for how close to C speed your PyPy benchmark results are. Cython is the language that many Python users write numeric calculations in, so the example you give would have been predestined for a Cython implementation – unless your real fear is the competition. 😉
FYI “Minima” is already plural (of minimum). “Minimas” is just wrong.
Fixed, thanks Chris.
What-about-comparison-2 with Shed Skin? Just-for-reference-2.
Nice writeup! I would be interested in the memory benchmarks as well.
Hello Carl, thanks for commenting =)
I’ll make another benchmark soon I get more time, so I’ll check memory footprint too.
Cool. No hurries though 🙂
Interesting results, thanks for taking the time to do this! Just a question about repeatability here…did you run the exact same code with the same random seed for all these tests?
Hello Brendan, that’s impossible since Jython has a wrapper for the Java prng, so they will never be the same evolution. But this is amortized due to the long run time.
If that’s the case, then it might be just dumb luck that one performs better/worse than the other (since you’ve only done three trials for each case). I suppose it’s not really feasible to get a good statistical sample (30+ trials for each case) due to the enormous run times, but do you think you could decrease the run time by decreasing the dimensionality of the Rastrigin function so that we could get statistically significant results?
As I have said later, it is amortized by the long run time, you run it and check by yourself.
If I decrease the dimensionality of Rastrigin, we’ll get still less feasible results. Different seeds only impacts on the result and in a subtle time margin, all testes were performed over 10k generations, no matter what results they give (although the same optimization results of the Rastrigin function were obtained).
“all testes were performed over 10k generations”
Ah, there’s the key. I missed that in your post and assumed you were running the optimization until it hit the global minimum, rather than for a set number of generations. So yes, I agree with you now.
Christian,
I’m using Pypy 1.4.1, and I’ve tried running a pyevolve GP on 3 different machines with different processors (AMD, Intel, 64 and 32 bit), and each run is much slower than regular python. The version of PyPy I’m running is the JIT version compiled version you can download on the pypy web site.
Are there special command line parameters I should be passing to pypy?
What command line arguments did you use to start and run your pypy scripts?
When I run a pystone test, pypy out performs the python version. So this really surprises me.
Thanks
Billy
To let you know, I’m running pyevolve 0.6, Multiprocessing off. A sample run of 10 generations of a GP (depth = 5) takes about 7 minutes on pypy, and only 2 minutes on python 2.7. pypy is jit enabled. Pysco is not being used.
I’ll try running a few other apps through pypy and python and see if the performance is about the same or if pypy is faster.
Any info you can provide will be helpful! 🙂
By the way, pyevolve is awesome!
Hello Billy, this is surprise to me too, I haven’t tested the GP core using PyPy yet, but it should not so slow as you reported. Have you tried to compare the CPython vs yours PyPy version with some Genetic Algorithm ? I’ll probably need to profile it in PyPy and check why is it so slow when compared to CPython. Have you checked the memory footprint between two interpreters ? What is the size of your population ?
Christian,
Eric Floehr in his pycon 2011 presentation states that pypy runs pyevolve code 3-4 times faster than cpython. But I’ve downloaded his code and ran it myself, and cpython runs the code 3 times faster than pypy (the exact opposite of what Eric proclaims, but I’m running python 2.6 (and 2.7) and pypy 1.4.1, so that might make some of the difference).
I’ve ran several of the examples in the trunk version of pyevolve, and for quick examples, cpython wins, and for longer jobs pypy usually wins (which is expected). I’ve also tried different population sizes for GPs to see if that would make a difference since memory usage would be different, and that does not affect the results when comparing cpython and pypy.
I’ve only ran Eric’s code on one machine, but I’ll try others and see if I get different results. If I can’t reproduce Eric’s results, I might send him an email and see if he can give me some details.
Again, I want to thank you for pyevolve, I’ve really enjoyed using it.
Billy
Would quite like to see Shedskin compared also.