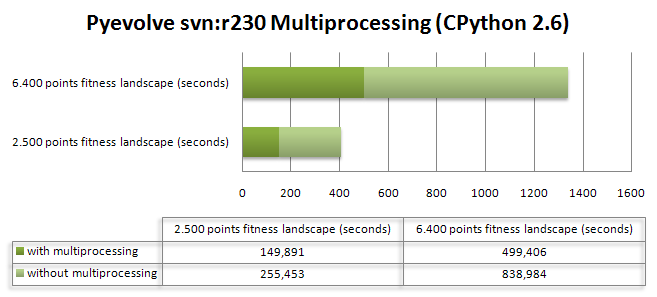
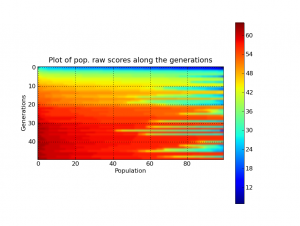
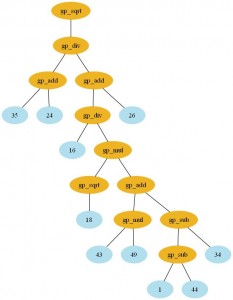
As we know, Genetic Programming usually requires intensive processing power for the fitness functions and tree manipulations (in crossover operations), and this fact can be a huge problem when using a pure Python approach like Pyevolve. So, to overcome this situation, I’ve used the Python multiprocessing features to implement a parallel fitness evaluation approach in Pyevolve and I was surprised by the super linear speedup I got for a cpu bound fitness function used to do the symbolic regression of the Pythagoras theorem: . I’ve used the same seed for the GP, so it has consumed nearly the same cpu resources for both test categories. Here are the results I obtained:
The first fitness landscape I’ve used had 2.500 points and the later had a fitness landscape of 6.400 points, here is the source code I’ve used (you just need to turn on the multiprocessing option using the setMultiProcessing method, so Pyevolve will use multiprocessing when you have more than one single core, you can enable the logging feature to check what’s going on behind the scenes):
from pyevolve import *
import math
rmse_accum = Util.ErrorAccumulator()
def gp_add(a, b): return a+b
def gp_sub(a, b): return a-b
def gp_mul(a, b): return a*b
def gp_sqrt(a): return math.sqrt(abs(a))
def eval_func(chromosome):
global rmse_accum
rmse_accum.reset()
code_comp = chromosome.getCompiledCode()
for a in xrange(0, 80):
for b in xrange(0, 80):
evaluated = eval(code_comp)
target = math.sqrt((a*a)+(b*b))
rmse_accum += (target, evaluated)
return rmse_accum.getRMSE()
def main_run():
genome = GTree.GTreeGP()
genome.setParams(max_depth=4, method="ramped")
genome.evaluator += eval_func
genome.mutator.set(Mutators.GTreeGPMutatorSubtree)
ga = GSimpleGA.GSimpleGA(genome, seed=666)
ga.setParams(gp_terminals = ['a', 'b'],
gp_function_prefix = "gp")
ga.setMinimax(Consts.minimaxType["minimize"])
ga.setGenerations(20)
ga.setCrossoverRate(1.0)
ga.setMutationRate(0.08)
ga.setPopulationSize(800)
ga.setMultiProcessing(True)
ga(freq_stats=5)
best = ga.bestIndividual()
if __name__ == "__main__":
main_run()
As you can see, the population size was 800 individuals with a 8% mutation rate and a 100% crossover rate for a simple 20 generations evolution. Of course you don’t need so many points in the fitness landscape, I’ve used 2.500+ points to create a cpu intensive fitness function, otherwise, the speedup can be less than 1.0 due the communication overhead between the processes. For the first case (2.500 points fitness landscape) I’ve got a 3.33x speedup and for the last case (6.400 points fitness landscape) I’ve got a 3.28x speedup. The tests were executed in a 2 cores pc (Intel Core 2 Duo).